SEA Phages
Dr. Sarah Ball has been involved with our Science Education Alliance (SEA) Phages CURE since its inception at OSU in 2011. Together, with Dr. Caroline Breitenberger, and Dr. Charles Daniels, SEA Phages lab began with an original cohort of 25 students. A part of some sections of Biology 1113, hundreds of students have participated in this CURE since that initial cohort. A national project, numerous universities and colleges across the United States are participating in this Phage-hunting.
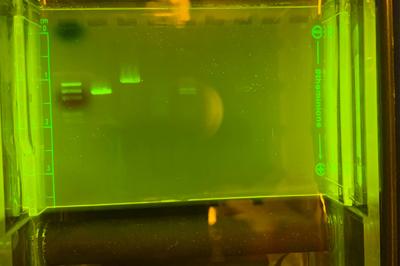
In this CURE, students isolate bacteriophages (viruses that infect bacteria) from soil samples collected from locations around campus, in Columbus, and beyond! Students purify their phage populations, and along the way learn important lab techniques, such as direct plating, and 3-phase streaks. Students then amplify the isolated sample of interest, and isolate DNA from the phage. They also conduct gel electrophoresis to characterize the bacteriophages.
Because this is a national project, information about the student isolated phages is added to a national database, called Phages DB. All of the phages isolated in Biology 1113 SEA Phages lab at OSU can be found on the OSU page of Phages DB. The DNA is then sequenced for a few of the phages isolated that semester. As of December 2021, 188 phages have been contributed to Phage DB.
After completion of the phage hunter CURE, students can enroll in Biology 2200 in a subsequent semester. In this course, the sequenced genome from a phage isolated the previous semester is annotated by students using DNAMaster and 51 completed genomes have been submitted to GenBank with 14 different clusters represented. The following list identifies phages sequenced and annotated by OSU undergraduate students.
Arthrobacter phage Xenomorph, complete genome Accession: MK919473.1
Arthrobacter phage EstebanJulior, complete genome Accession: MK919476.1
Arthrobacter phage Grekaycon, complete genome Accession: MK919479.1
Arthrobacter phage DreamTeam, complete genome Accession: MK919484.1
Arthrobacter phage Kumotta, complete genome Accession: MK977703.1
Arthrobacter phage Adat, complete genome Accession: NC_042020.1
Arthrobacter phage Breylor17, complete genome Accession: MH450115.1
Mycobacterium phage Buckeye, complete genome Accession: MH450116.1
Arthrobacter phage Daiboju, complete genome Accession: MH450117.1
Arthrobacter phage KeaneyLin, complete genome Accession: MH450120.1
Arthrobacter phage KingBob, complete genome Accession: MH450121.1
Mycobacterium phage KingTut, complete genome Accession: MH450122.1
Arthrobacter phage MargaretKali, complete genome Accession: MH450123.1
Arthrobacter phage MediumFry, complete genome Accession: MH450125.1
Mycobacterium phage GeneCoco, complete genome Accession: MH479916.1
Arthrobacter phage Synepsis, complete genome Accession: MH479926.1
Mycobacterium phage MikeLiesIn, complete genome Accession: MH051266.1
Arthrobacter phage GurgleFerb, complete genome Accession: MF668273.1
Arthrobacter phage Nellie, complete genome Accession: MF668279.1
Arthrobacter phage Temper16, complete genome Accession: MF668285.1
Arthrobacter phage Niktson, complete genome Accession: MF038790.1
Arthrobacter phage ElephantMan, complete genome Accession: MF038791.1
Mycobacterium phage WillSterrel, complete genome Accession: KX576644.1
Mycobacterium phage Derpp, complete genome Accession: KX576645.1
Mycobacterium phage TyrionL, complete genome Accession: KX576646.1
Mycobacterium phage CharlieGBrown, complete genome Accession: KX576647.1
Mycobacterium phage Tasp14, complete genome Accession: NC_028784.1
Mycobacterium phage Ariel, complete genome Accession: NC_028876.2
Mycobacterium phage Squirty, complete genome Accession: NC_026588.1
Mycobacterium phage BuzzLyseyear, complete genome Accession: NC_026593.1
Mycobacterium phage Vivaldi, complete genome Accession: KM347890.1
Mycobacterium phage Murphy, complete genome Accession: NC_021305.1
As part of this course, 2-3 students a semester are selected to present results at an HHMI symposium in Northern Virginia. The poster submissions, with undergraduate students in bold are listed below.
S. Randall, S. Ball, C. Breitenberger, C. Daniels (2019) Singleton No More: MargaretKali Finds a Match in Kumotta. The 11th Annual SEA PHAGES Symposium. HHMI Janelia Farms Research Complex, Ashburn, VA
N. Cervelli, D. Najjar, S. Ball, C. Breitenberger, C. Daniels (2018) Genomic Analysis of Arthrobacteriophage MargaretKali, a Singleton. The 10th Annual SEA PHAGES Symposium. HHMI Janelia Farms Research Complex, Ashburn, VA
C. Bertolini, A. Tyransky, C. Breitenberger, S. Ball, C. Daniels (2017) Comparative Analysis of Cluster AV Arthrobacteriophages, Including Recent Isolates Adat, Gurgleferb, and Nellie. The 9th Annual SEA PHAGES Symposium. HHMI Janelia Farms Research Complex, Ashburn, VA
S. Hastings, A, Moore, S. Ball, C. Breitenberger (2016) Sojourns in the F Cluster. The 8th Annual SEA PHAGES Symposium. HHMI Janelia Farms Research Complex, Ashburn, VA
G. Brockman, E. Lemanski, S. Ball, C. Breitenberger, C. Daniels, L. Webb (2015) Four years of mycobacteriophage isolation at Ohio State. The 67th Annual SEA PHAGES Symposium. HHMI Janelia Farms Research Complex, Ashburn, VA
A.J. Brusnahan, M. D. Postolowski, S. Ball, C. Breitenberger, and C. Daniels (2014) Comparative analysis of F Cluster mycobacteriophages, including recent isolates Buzzlyseyear and Squirty. The 6th Annual SEA PHAGES Symposium. HHMI Janelia Farms Research Complex, Ashburn, VA
C. Akusoba, Z. Al-Firdaus, N. Arora, R. Bari, J. Brown, I. Chancellor, A. Gajjala, J. Galletta, H. Goldsmith, J. Gonzalez, E. Gough, S. Gregory, J. Guo, B. Harbaugh, N. Hegarty, N. Justus, B. Kaur, E. Kittaka, R. Klein, E. Martin, L. McNulty, H. Mehalik, S. Morin, S. Mosteller, Z. Murphy, H. Nootbaar, I. Nurko, E. Osborne, R. Pedagandham, F. Qureshi, M. Reilly, E. Reinhart, K. Roush, T. Schmidt, K. Schroeder, M. Sherer, M. Shipton, K. Shively, C. Siesel, M. Tounkara, S. Warwar, L. Webb, D. Winders,S. Ball, C. Breitenberger, and C. Daniels (2013) Mycobacteriophage Squirty defines a new subcluster, F3. The 5th Annual SEA PHAGES Symposium. HHMI Janelia Farms Research Complex, Ashburn, VA
M. Anderson, N. Justus, L. Webb, R. Bari, J. Bir, P. Consiglio, E. Domingo, N. Elfessi, S. Frankel, G. Hwang, T. Lang, Y. Lu, M. Mollica, J. Morena, S. Petrofski, X. Rui, M. Shafiq, T. Varughese, S. Ball, C. Breitenberger, and C. Daniels (2012) Isolation and Genomic Sequencing of the Novel Mycobacterial Phage, Murphy. The 4th Annual SEA PHAGES Symposium. HHMI Janelia Farms Research Complex, Ashburn, VA
Interestingly, the types of bacteria hosts are varied. So far, students in Biology 1113 have isolated bacteriophages from Arthrobacter sp., Mycobacterium smegmatis, and Gordonia terrae. Phages representing 12 different clusters have been sequenced giving us just a small glimpse at the incredible diversity waiting to be uncovered. There are always more interesting phages to be discovered!
We have also developed a helper program in which students who successfully complete the course, and its associated genome course become student assistants in the classroom. These helpers have been invaluable assistants to our Teaching Assistants. Other undergraduate students have been instrumental on our lab prep team, making sure that students in the course have the supplies they need.
